Researchers at the Massachusetts Institute of Technology (MIT) and Georgia Tech have automated the process of finding and recording information from cells in living brains, as reported in the May 6 issue of Nature Methods.
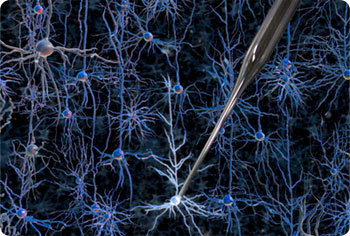
“Autopatching makes recording the electrical and circuit properties [of cells] straightforward and effortless for the operator,” Craig Forest, assistant professor at Georgia Tech and one of the members of the research team, told TechNewsWorld.
This technology “can be integrated with molecular analysis, such as gene expression measurement, and cell staining for shape, or morphology, measurement. Together, electrophysiology, gene expression and morphology comprise a complete description for healthy cells and cells that are under the influence of drugs or diseased for comparison,” he said.
“For example, such a study could be used to discover the cellular processesunderlying diseases like epilepsy, narcolepsy, Alzheimer’s … or fordrugs like anesthesia,” he added.
The Story Behind the Idea
It’s been possible for about 30 years to look at individual brain cells and find out what their electrical and circuit properties are. The technique is called “whole-cell patch clamping.” It was developed in the late 1970s and early 1980s by Erwin Neher and Bert Sakmann, who received the Nobel Prize in Physiology or Medicine in 1991 for their work.
In whole-cell patch clamping, scientists essentially lower a tiny glass pipette onto a cell, apply suction to make sure the two are sealed together tightly, and push a silver wire in the pipette into the cell to detect its electrical signals. That signal’s fed back to amplifiers.
The technique is most useful when it comes to excitable cells such as neurons and muscle fibers.
However, using the whole cell patch clamp technique requires extensive training and even highly trained scientists can examine data from only a few cells a day.
That led Suhasa Kodandaramaiah to work with Forest and Ed Boyden, an associate professor at MIT, and others on a way to automate the process.
Playing With the Brain
Kodandaramaiah found that the whole-cell patch clamp process of lowering the pipette can be reduced to a reliable four-stage algorithm.
The researchers use a piezoelectric stepper motor driving a precision linear stage. A computer running the control algorithm sends signals to the motor. Once the robot arm gets to the area of interest, it takes small steps of 1-2 micrometers until the micropipette, which has a tip 1 micrometer in diameter, touches a neuron, Forest said. Neurons have a diameter of 10 micrometers.
The robot takes three to seven minutes to perform the entire process. It detects cells by analyzing the sequence of changes in electrical impedance over time. It measures impedance 10 times a second.
The researchers demonstrated the robot’s use on live mice.
Getting Better All the Time
The robot can detect cells with 90 percent accuracy and establish a connection with detected cells about 40 percent of the time.
Since it’s going to be poking around in a live brain, those figures might seem a tad low. However, “The success rate of the best humans is about 20 percent,” Forest said. “The best humans can do three to 10 neurons a day.”
The real advantage of automation is that “the process is effortless and repeatable and systematic, which makes it easily adopted by others, and easily integrated with other technologies,” Forest pointed out.
Breaking New Ground
Commercial automated whole-cell patch clamp systems have been around for “quite a while,” Gary Cheng, senior application scientist at ALA Scientific Instruments, told TechNewsWorld.
One such line of systems, the IonFlux family from Fluxion Biosciences, is priced at “about (US)$100,000 to $300,000,” Tom Driscoll, a vice president at the company, told TechNewsWorld. The larger ones are used by pharmaceutical and biotech companies.
However, “other systems for patch clamping either work on neurons in suspension or cultured neurons or slices of neurons in vitro because this is much easier,” Forest said.
The MIT-Georgia Tech robotic arm can be used in vivo, meaning on living brains.
“Neuroscientists would prefer to work in vivo where the neuron is communicating with its neighbors in the way they were originally supposed to, with the original synapses and the internal life support system the body provides,” Forest stated